
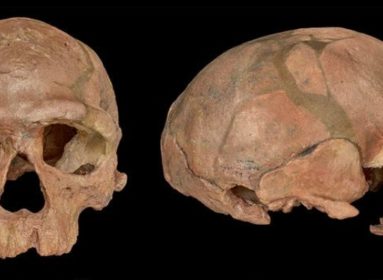
Durvasula & Sankararaman provide complementary lines of evidence for archaic introgression into four West African populations. Image credit: Bob Wilder, University at Buffalo.
Four West African populations derive 2 to 19% of their genetic ancestry from a yet-undiscovered species of archaic hominin that diverged before the split of modern humans and the ancestors of Neanderthals and Denisovans.
Contemporary people who have ancestors from Europe, Asia, and Oceania carry DNA from two archaic species, Neanderthals and Denisovans, making up 1-4% of their genome.
These genetic segments arrived in modern humans through introgression, the process by which members of two populations mate, and the resulting hybrid individuals then breed with members of the parent populations.
Recent studies have shown that, though modern West Africans do not have Neanderthal or Denisovan ancestry, there may have been introgression by other ancient hominins in their past.
In a new study, University of California, Los Angeles researchers Arun Durvasula and Sriram Sankararaman compared Neanderthal and Denisovan DNA with genomes of 405 individuals from West Africa.
The scientists focused on four contemporary West African populations: Yoruba from Ibadan, Esan from Nigeria, Mende from Sierra Leone, and Gambian.
They found differences that could be best explained by introgression by an unknown archaic hominin whose ancestors split off from the human family tree before Neanderthals.
The data suggest this introgression may have happened relatively recently, or it may have involved multiple populations of archaic human, hinting at complex and long-lived interactions between anatomically modern humans and various populations of archaic hominins.
“Combining our results across the West African populations, we estimate that the archaic population split from the ancestor of Neanderthals and modern humans 360,000 years to 1.02 million years ago, and subsequently introgressed into the ancestors of present-day Africans 0-124,000 years ago, contributing 2 to 19% of their ancestry,” the authors said.
Dr. Durvasula and Dr. Sankararaman also examined the frequencies of archaic DNA segments to investigate whether natural selection could have shaped the distribution of archaic genetic variants.
“We found 33 loci with an archaic segment frequency of over 50% in Yoruba and 37 loci in Mende,” they said.
“Some of these genes are at high frequency across both Yoruba and Yoruba, including NF1 (a tumor suppressor gene), MTFR2 (a gene involved with mitochondrial aerobic respiration in the testis), HSD17B2 (a gene involved with hormone regulation), KCNIP4 (a gene involved with potassium channels), and TRPS1 TRPS1 (a gene associated with trichorhinophalangeal syndrome).”
“Three of these genes have been found in previous scans for positive selection in Yoruba: NF1, KCNIP4, and TRPS1. On the other hand, we do not find elevated frequencies at MUC7, a gene previously found to harbor signatures of archaic introgression.”

The team calls for more analysis of modern and ancient African genomes to reveal the nature of this complex history.
“The signals of introgression in the West African populations that we have analyzed raise questions regarding the identity of the archaic hominin and its interactions with the modern human populations in Africa,” the researchers said.
“A detailed understanding of archaic introgression and its role in adapting to diverse environmental conditions will require analysis of genomes from extant and ancient genomes across the geographic range of Africa.”
Source: Enrico de Lazaro / Science Advances.


















































Comments are closed.